Raman spectroscopy is a scattering
technique that relies on the inelastic scattering of photons. The excitation of
molecular bonds results in the shifts of the energy of laser photons and thus
frequency change of the incident light. Therefore, a compound can have a
specific Raman spectrum, while a Single-cell Raman Spectrum (SCRS) can depict
the overall profile of metabolites in the cell, i.e., the metabolic state of
the cell at a particular instance.
We have proposed the
"Ramanome" concept, which is the collection of the SCRS from a
cellular population (or a consortium, for “Meta-ramanome”) at a given
instance. Each SCRS can have over 1500 Raman peaks, while each peak or
combination of peaks can potentially represent a metabolic phenotype, therefore
a ramanome data point would capture the single-cell-resolution metabolic
phenome of the system at a certain state. Notably, within a ramanome, even
though the genomes of the cells are identical, the SCRS of the cells are
distinct; such intercellular heterogeneity of SCRS is an inherent feature of
cellular systems, and is critical for single-cell analysis and interpretion.
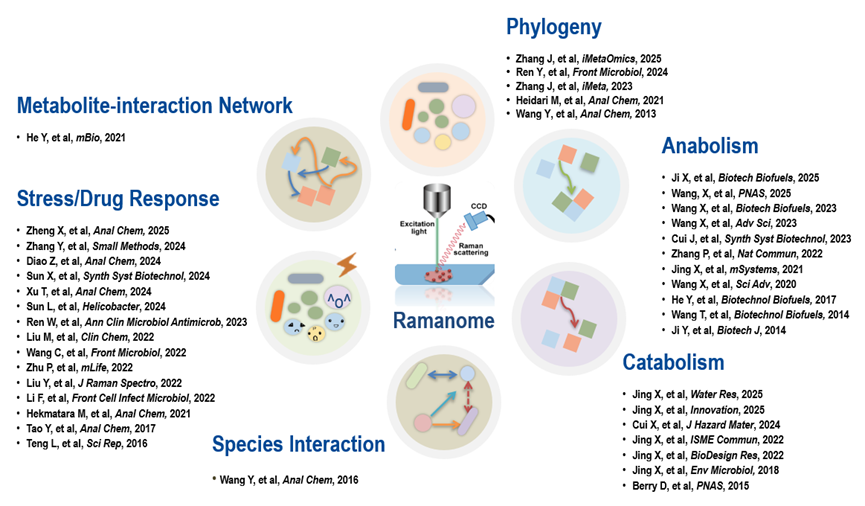
We together with collaborators have demonstrated that
ramanome can be employed to quantitatively profile a wide variety of metabolic
phenotypes for individual cells, such as quantifying the intake rate of hydrogen- and carbon-containing substrates, determining the
diversity and content of various Raman-sensitive intracellular products (pigments, triglycerides, starch, proteins, etc.),
characterizing the environmental stress responses of cells (e.g., antimicrobial susceptibility of pathogens, mechanisms of microbial drug response, drug resistance and its mechanisms for tumor
cells, etc.), detecting intercellular metabolism interactions, reconstructing intracellular metabolite interconversion networks (Intra-Ramanome Correlation Analysis; IRCA), and
distinguishing different microbial (or microalgal) species.
The application of ramanome is rapidly expanding.
Ramanome has broad applications. For example, to meet
the challenge in precision medicine, we invented the D2O-Ramanometry technology to rapidly measure stress
response at single-cell level, and proposed the concept of MIC-MA (the lowest drug dose at which the metabolic activity of
"every" cell is "completely" inhibited). As an innovative
parameter to quantify drug sensitivity, MIC-MA is faster, more accurate, deeper
in revealing mechanism, broader in application and easier in measurement than
the traditional MIC. Then, we have developed the first Clinical Antimicrobial
Susceptibility Test Ramanometry (CAST-R ), and established the "Clinical Demonstration
Network for Rapid AST at Single-cell level" together with clinical
partners throughout China. MIC-MA/CAST-R has demonstrated its advantages in the
treatment of acute bloodstream infections (tigecycline, etc.), chronic infections (Helicobacter pylori, etc.), oral disinfectant efficacy evaluation and
screening (for bacteria and fungi), and highly sensitive drug resistance phenotyping of
mutants. This principle is also applicable to the
evaluation and screening of metabolic activity and/or resistance for industrial/environmental microorganisms.
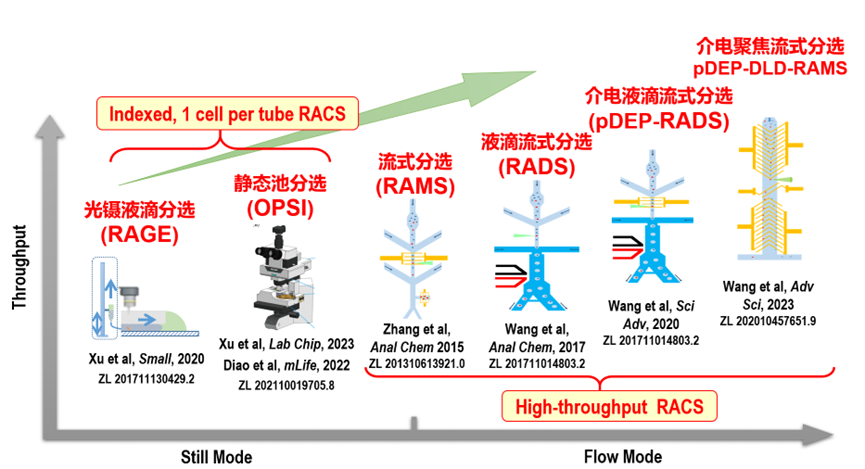
In order to obtain the single cells of
targeted metabolic phenome from a cellular population or consortium (for e.g.,
single-cell sequencing or culture), we have developed and commercialized a
series of Raman-guided single-cell sorting devices. These Raman-activated Cell
Sorting (RACS) technologies are broadly classified into
two categories: stationary and microflow, depending on the motion state of
cells at the time of Raman acquisition.
The former includes Raman tweezers,
Raman-Activated Cell Ejection (RACE and its upgraded version), and Raman-Activated Gravity-driven single-cell
Encapsulation (RAGE), etc. This category of RACS devices
mainly focuses on precise and indexed Raman-based single-cell collection and
sorting. The latter, capable of sorting cells in a high-throughput manner,
includes Raman-activated Microfluidic Sorting (RAMS), Raman-activated Droplet Sorting (RADS), and positive dielectrophoresis based
Raman-activated Droplet Sorting (pDEP-RADS). In addition, we have introduced a number of
downstream single-cell sequencing or culture technologies that are directly
coupled to RACS, such as precisely-one-bacterial-cell high-coverage genome
sequencing (RAGE-Seq),
single-cell culture (RACS-Culture), and
Addressable Dynamic Droplet Array (aDDA)-based single-cell sequencing technologies.
For Ramanome data analysis, we have
proposed novel algorithms and concepts, such as Intra-Ramanome Correlation
Analysis (IRCA) and Raman Barcode of Cellular-response to
Stresses (RBCS). We also developed the Ramanome Ensemble Learning algorithm and published
the first Micro-Algal Ramanome Database (MARD). These algorithms, software and databases are all
generally applicable.
With the generous support of the National
Natural Science Foundation of China, Ministry of Science and Technology of
China, and Chinese Academy of Sciences, we have developed a series of
Ramanome/RACS-based instruments, which have entered the market, such as
Clinical Antimicrobial Susceptibility Test Ramanometry (CAST-R), Flow-mode Raman-activated Cell Sorter (FlowRACS), Raman-activated Single-Cell Sorting and
Sequencing (RACS-Seq),
Single-cell Microdroplet Sorting System (EasySort Lego / Compact), etc.
Using these original
instruments, we have established a platform to support the entire workflow that
links single-cell metabolic phenome profiling and the corresponding
high-quality single-cell genome sequencing. Moreover, our instruments have been
employed in many research areas, such as high-throughput screening of enzyme activity in
vivo, rapid profiling of metabolic functions
in cellular factories,
high-throughput screening of metabolic phenotypes of mutants (oil-producing super microalgae induced by blue light), and reconstruction of metabolite-conversion
networks. For microbiome samples, using RACS-Seq which
is equipped with the RAGE chip, we have, for the first time, demonstrated the ability to
simultaneously obtain metabolic phenome and high-coverage genome from precisely
one bacterial cell, from both clinic samples (e.g., urine, gastric mucosa, etc.) and
complex environmental samples (e.g., soil, ocean, etc.).
In
addition, we established a new technique called single-cell Raman-activated
Cell Sorting and Cultivation (scRACS-Culture), for metabolic-function based assessment and mining of live probiotics
directly from complex environmental samples (such as mining in
situ phosphate solubilizing microbes from urban wastewater),
which are advantageous because it is of single-cell precision, culture-free,
and does not require fluorescence probes. This "screen first, culture
second" approach is a new route for functional mining of microbiomes.
By quantitatively converting the molecular
spectrum of intracellular metabolites into key metabolic phenotypes such as
substrate intake, product synthesis, response to environment and metabolite
conversion, ramanome is a kind of “omics” that is closest to “function”,
and a more direct way to probe the metabolism of cells. Moreover, ramanome-based tools
offer many advantages, such as label-freeness, non-destructiveness,
landscape-like phenotyping, high-speed, low-cost, automation, general
applicability to all cell types, and ability to couple to downstream
single-cell sequencing, mass spectrometry or cell culture. Thus ramanome is
highly complementary to those existing omics tools, and together they formulate
a complete single-cell multi-omics platform.
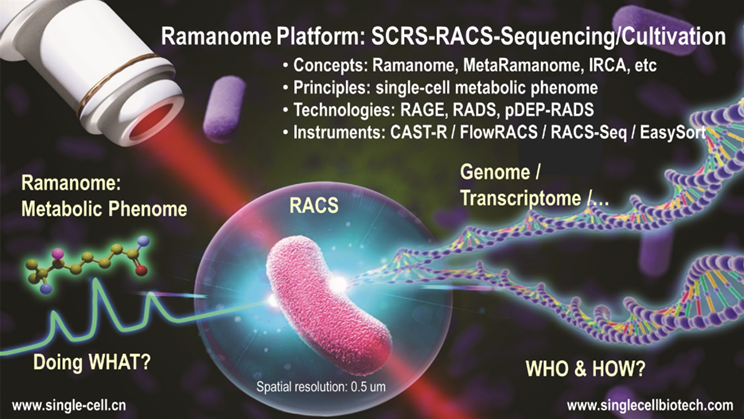
In summary, Ramanome Technology Platform (RTP), which establishes
the link between metabolic phenome and genome (or transcriptome, proteome, etc)
at the precision of one cell (or one organelle for an eukaryotic cell), answers
the fundamental question of "Who is doing What, and How" at the deepest
level. Therefore, RTP has become a powerful tool for functional mining and
mechanistic dissection of human, animal, plant and microbial single-cells, and
is serving a wide range of fields such as precision medicine, synthetic
biology, one-health, biosecurity, and ecological surveillance.